Efficient Storage of Images onto DNA Using Vector Quantization
Résumé
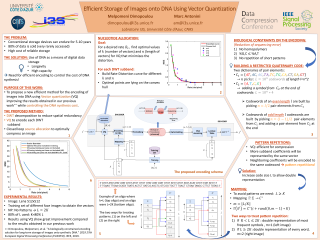
The archiving of digital data is becoming very challenging as conventional electronic devices wear out in time leaving at stake any data that has been stored in them. Therefore, data migration is necessary every 5-10 years. Unfortunately, the maintenance and replacement of servers in big data centers is very expensive both in terms of money and energy. DNA data storage is a new evolving technique which proposes an efficient and eco-friendly solution while also promising data longevity due to the use of DNA as a means of digital data storage. Storing digital information into DNA is feasible by encoding it in a quaternary representation using the four DNA nucleotides (nts) A, T, C and G which is later going to be synthesized into DNA. Then, thanks to some machines which are called sequencers, one can read back the synthetic DNA and retrieve the corresponding quaternary sequence. Finally, using a decoding procedure the stored data can be recovered. The most challenging part of this procedure is the fact that the reading process of sequencing is error-prone. Respecting some special restrictions in the encoding can lead to a more reliable reconstruction. More precisely the existence of homopolymers, a high percentage of G and C or the repetition of short patterns in the encoded sequence can lead to high se-quencing error. Furthermore, as DNA synthesis costs several dollars per DNA strand (200 nts), it is also important to ensure that the encoding is efficient in terms of data compression. Consequently the selection of an appropriate encoding algorithm is highly important. In this work we propose a new end-to-end encoding schema which is specific for the efficient storage of images onto synthetic DNA. This algorithm uses a DWT to create wavelet sub-bands and then quantizes each one of them using a Vector Quantizer(VQ). Quantization is optimized by a source allocation algorithm which allows selection of an optimal number of vectors and an optimal length of vectors that provide the minimum distortion at a given rate. The optimally compressed sub-band coefficients are then encoded into a sequence of A, T, C, G using a robust algorithm which respects the restrictions imposed by the biological procedure of DNA sequencing to ensure reliability of the decoding. The novelty of this work compared to the existing state of the art lies in the implementation of a full encoding workflow which minimizes the cost of storing images into DNA while avoiding the creation of patterns which can occur when using VQ. Our results show great improvement in the rate-distortion curve from our previous experiments that have been using uniform quantization while also producing DNA strands which respect all the necessary sequencing restrictions. More precisely VQ allows us to improve the coding rate to 3.34 bits/nt which is a very good performance compared to the state of the art.
Fichier principal
 Dimopoulou_DCC_poster.pdf (853.6 Ko)
Télécharger le fichier
Dimopoulou_summary_DCC_2020.pdf (60.01 Ko)
Télécharger le fichier
Dimopoulou_DCC_poster.pdf (853.6 Ko)
Télécharger le fichier
Dimopoulou_summary_DCC_2020.pdf (60.01 Ko)
Télécharger le fichier
Origine : Fichiers produits par l'(les) auteur(s)
Origine : Fichiers produits par l'(les) auteur(s)